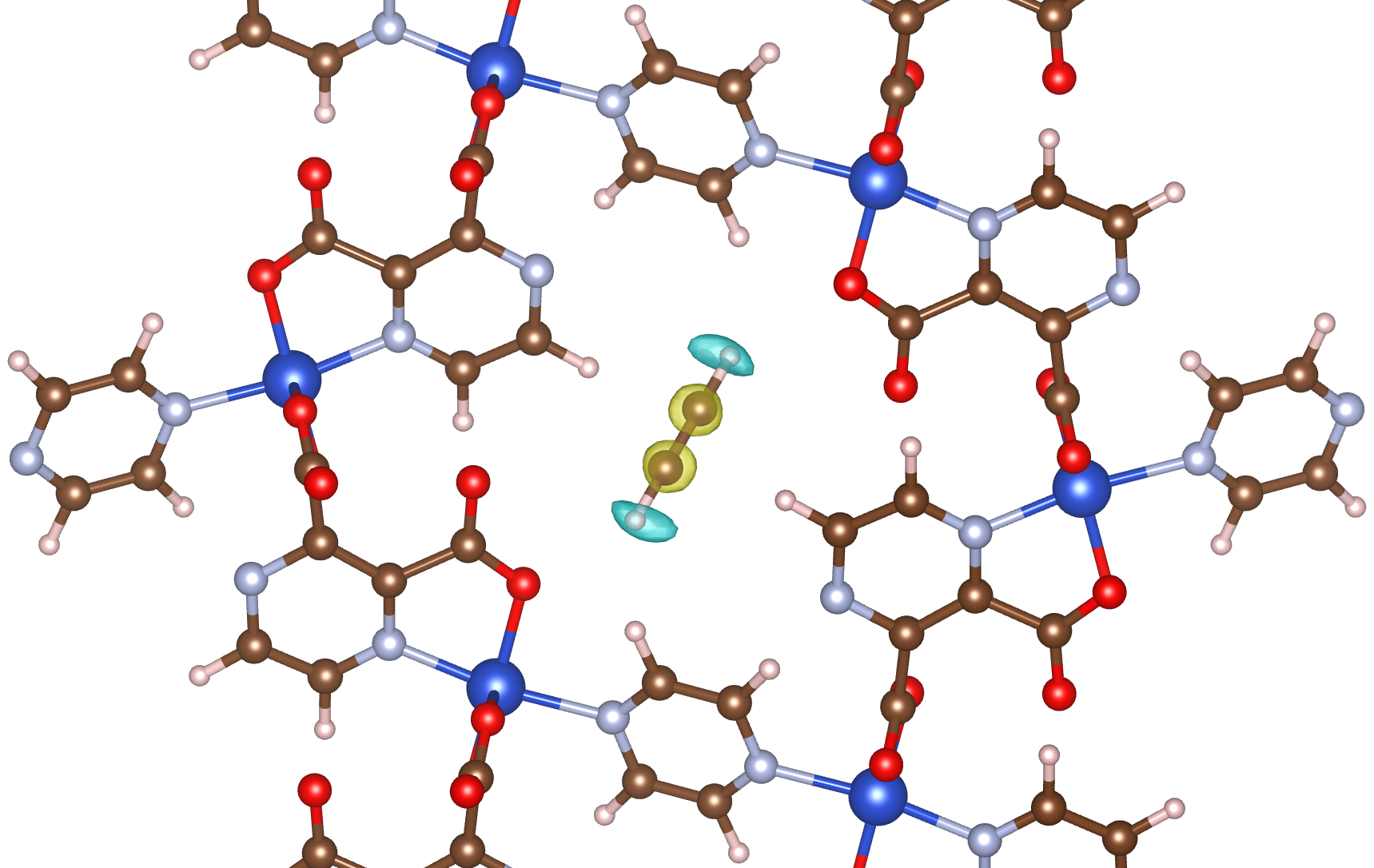
This section illustrates advanced density grid features in RASPA3, including site-resolved probability plots, equitable binning, and their combined use for analyzing adsorption behaviour and identifying binding regions.
Table of Contents
- Binary mixture and site-resolved density grids
1_mc_adsorption_binary_mixture_co2_ch4_in_irmof_1_split_density_by_site - Effect of equitable binning on density grids
2_mc_adsorption_of_methane_in_mfi_standard_binning3_mc_adsorption_of_methane_in_mfi_equitable_binning - Combined use: site-resolved grids, equitable binning, and binding site analysis
4_mc_adsorption_of_co2_in_calf_20_methyl_split_density_equitable_binning5_mc_adsorption_of_c2h2_in_cpl_1_split_density_equitable_binning6_mc_adsorption_of_co2_in_calf_20_split_density_equitable_binning
Binary mixture and site-resolved density grids
This example demonstrates the generation of site-resolved density grids for a binary mixture of CO2 and CH4 adsorbed in IRMOF-1. By restricting the density grid calculation to selected pseudo atom types, separate probability plots are written for each interaction site. This allows adsorption behaviour to be resolved at the atomic level in multi-component systems. In this example, density grids are generated only for the pseudo atoms associated with CO2. If density grids for CH4 interaction sites are also of interest, the corresponding pseudo atom name can be added to the "DensityGridPseudoAtomsList" to generate those probability plots as well.
Separate density grid files are written for each specified pseudo atom type and stored in the density_grids directory.
Effect of equitable binning on density grids
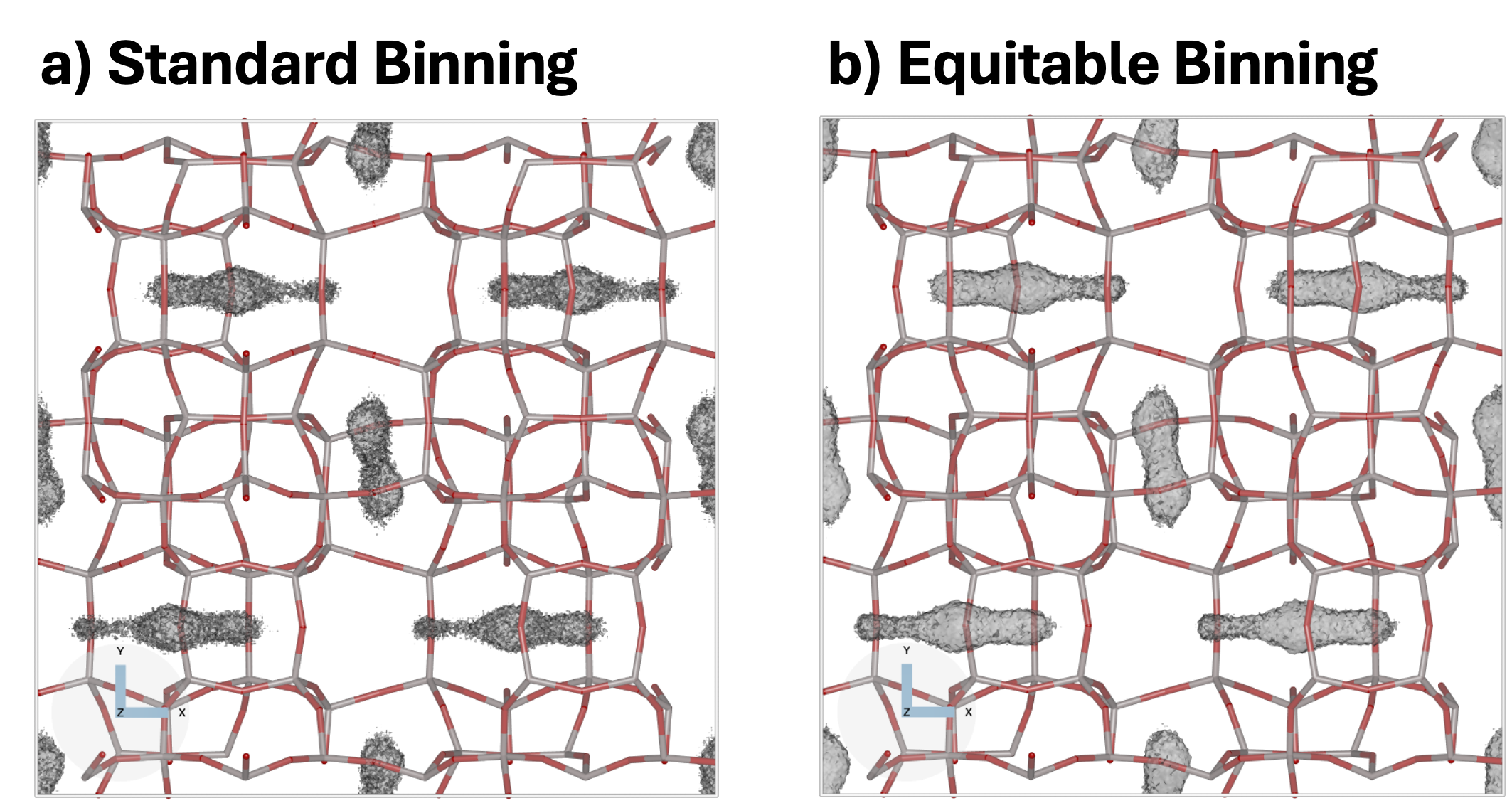
The following examples compare conventional histogram binning with equitable binning for methane adsorption in the MFI framework. Both simulations use identical settings except for the binning strategy.
Standard binning
Equitable binning
With equitable binning, each guest site contributes fractionally to neighbouring voxels based on its position, reducing discretization artefacts and producing smoother probability distributions. Neither standard nor equitable binning is inherently superior, and for well converged simulations the resulting local maxima are expected to be very similar. The figure below compares the two binning schemes at a representative isovalue, highlighting the primarily visual nature of the differences.
Combined use: site-resolved grids, equitable binning, and binding site analysis
This section demonstrates the combined use of site-resolved density grids and equitable binning for chemically complex systems. Examples include CO2 adsorption in CALF-20 (with and without methyl functionalization) and C2H2 adsorption in CPL-1.
The resulting density grids provide smooth, atom-specific probability distributions that highlight preferred adsorption regions within the frameworks.
Such probability plots can be further analyzed using external post-processing tools to identify high-probability regions and extract representative binding site configurations. The figures shown here were generated using external analysis routines to extract and fit binding site geometries directly from adsorbate probability distributions (APD) such as the RASPA-generated density grids. In these visualizations, the probability density highlights regions where a guest molecule is most likely to reside within the framework. For example, in the CPL-1 case, individual atomic contributions are resolved, with different colours used to represent distinct atom types within the guest molecule (e.g., carbon and hydrogen), providing an intuitive picture of both molecular orientation and preferred adsorption sites. The grid resolutions used for these examples were selected to correspond to a real-space grid spacing of 0.15 Å, and the supercell sizes were determined by the van der Waals cutoff. As a result, the displayed grids differ from the default [128, 128, 128] resolution and also differ between systems. These workflows operate independently of the simulation itself.